|
Early bird gets the wormDivers collect marine invertebrate to study genetic cold adaptationsPosted November 4, 2011
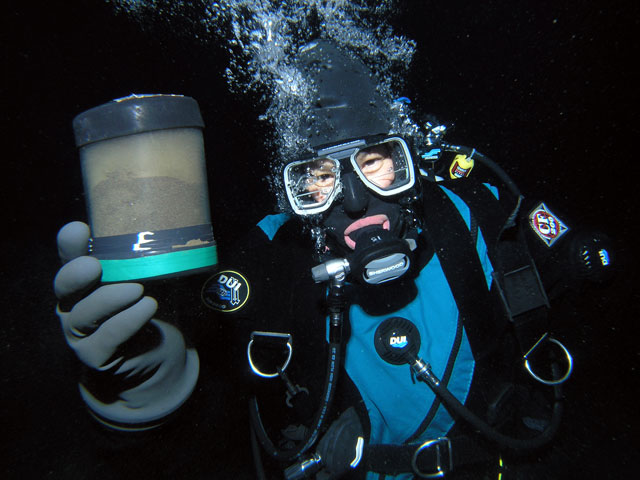
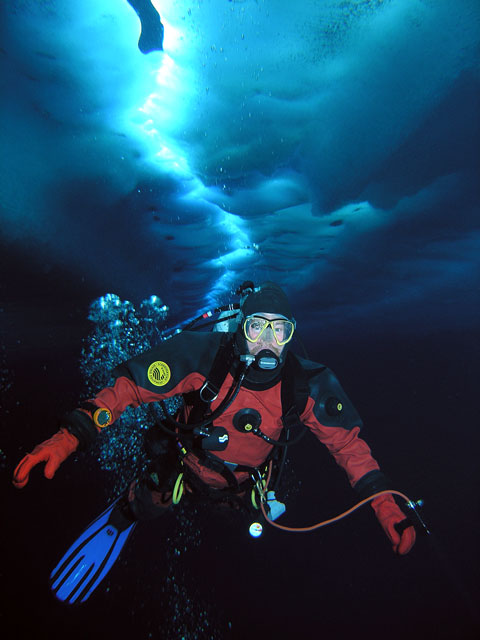
A little worm that inhabits organically enriched sediment in the subfreezing waters of Antarctica may help shed light on how all organisms interact with their environment on a genetic level. Adam Marsh However, it’s not really Capitella’s proclivity to putrid-smelling environments that has research divers digging through sediments on the seafloor, using a surface-supplied air system to protect them from the contaminated environment. Rather, Capitella perarmata is an attractive study subject because of a closely related temperate species called Capitella teleta. The latter is one of the few marine organisms for which a well-defined genome sequence exists, according to Marsh, associate professor of marine biosciences at the University of Delaware. This genetic sequence gives his team a leg up as it investigates how the polar environment specifically influences Capitella perarmata’s epigenome. 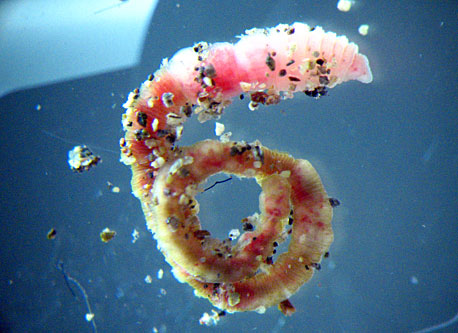
Photo Credit: Annamarie Pasqualone/PolarTREC
Capitella worms have adapted to the minus 1.8 degree Celsius water temperatures in Antarctica.
“It’s advantageous because there are a lot of genomic resources already available for this genus of marine species,” said Marsh, who has made 15 trips to the Antarctic. Previous work included investigating how critters like sea urchins and starfish have adapted to the cold environment of McMurdo Sound. Those experiments included using the sperm and eggs from collected specimens to create embryos to determine the genetic switches that control their growth rates. “This work we are currently pursuing is sort of the next level of what determines differences in global patterns of metabolism, global patterns of gene expression,” Marsh explained. The scientists are working in the emerging field of epigenetics, which translates to “above the genome.” In a well-used computer analogy, if the genome is an organism’s hardware, the epigenome is the software that tells the computer when to work, how to work and how much work to do. Tiny chemical switches in the form of methyl groups can shut genes down or turn genes on. Research has shown that epigenetic changes accumulate throughout an organism’s lifetime, based on environmental stimuli, and that those changes can be inherited in the next generation. “We already have an idea of what the temperate worm genome and epigenetic profiles look like, so by looking at the polar species, those polar differences should be statistically more evident in comparisons that we make,” Marsh said. “What we find via this one worm, Capitella, in terms of temperature adaptation, should be very applicable to other invertebrate species here as well.” This is the project’s second and final field season. Last year, the team had trouble finding Capitella in any abundance and settled on working on a different species. However, this season, Marsh recruited Stacy Kim “This is a way of combining lab techniques with field techniques, which is something that we’re both interested in doing,” Kim said. For example, in the future, the team is interested in investigating the epigenetic patterns of gene expression for other cold-adapted organisms as they apply to feeding strategies, particularly those critters that live on the seafloor, which is often scoured clean by icebergs. Imagine living with the specter of a wrecking ball obliterating your kitchen every so often. What strategies would you adapt to deal with such environmental stress? “We’re trying to coordinate with what we find on this project via epigenetic patterning to down-regulate metabolic shifts in organisms, into a stronger field context of flow disturbance,” Marsh said. Most of the genomic lab work will take place back in the United States. While in McMurdo, University of Delaware graduate students Stephanie Guida and Annamarie Pasqualone extract the marine worms from the mud and transfer them to fresh sediment. The animals will be cultured in the McMurdo lab for experiments mostly involving metabolic rates. The scientists will measure metabolism at different elevated temperatures. “The elevated temperature is a tweak, in a sense, to find those locations where [DNA] methylation does shift as a function of temperature,” Marsh said. Capitella’s temperate cousin will undergo similar experiments. “We will compare the temperate species and polar species in terms of how they respond to temperature to make that correlation about which sites, what kind of genes, are more likely involved in that regulatory shift,” Marsh said. Of course, the project is about more than simple marine worms. The epigenetic mechanisms at work in Capitella are the same ones that can possibly tag a human cell to turn cancerous or cue the onset of a degenerative disease. Marsh said his lab is collaborating with a group at the Pennsylvania School of Medicine that is interested in Parkinson’s disease by studying degenerative nerve conditions in mice. “The kinds of work that we’re doing here on the Capitella worm, we’re already drawing parallel experiments now in a mouse system to try and see if epigenetic patterning could be active in other sorts of nerve degenerative diseases,” he explained. “Having a successful epigenetic project with Capitella just opens the door for justifying working with any species, so it really is worthwhile collecting all of that sequencing information.” NSF-funded research in this story: Adam Marsh, University of Delaware, Award No. 0944557 |



For USAP Participants |
For The Public |
For Researchers and EducatorsContact UsNational Science FoundationOffice of Polar Programs Geosciences Directorate 2415 Eisenhower Avenue, Suite W7100 Alexandria, VA 22314 Sign up for the NSF Office of Polar Programs newsletter and events. Feedback Form |





